I’m starting to come around on voltage imaging. I haven’t been a fan of it for a number of reasons.
- The response sizes suck. Classic dyes and genetically encoded systems get a few percent fluorescence change at best.
- The response speeds suck. Measuring continuous current injections from -100mV to +150mV is not very interesting. Action potentials are interesting. But they are fast.
- Toxicity. The dyes kill neurons, or strongly perturb their electrical properties.
OK, voltage-sensitive imaging isn’t totally useless, for example see Carl Petersen’s recent paper on Spatiotemporal Dynamics of Cortical Sensorimotor Integration in Behaving Mice (2007). But if the above problems could be solved, then voltage sensitive imaging would be a strong competitor to calcium imaging for the non-invasive, high-resolution monitoring of patterns of network activity. There has been considerable progress ameliorating these problems in the past few years, much of it by a consortium of labs (Isacoff, Knöpfel, Bezanilla, Miesenböck, and others) focused on these issues. (Umlaut’s apparently help in this field).
First, let’s look at a minor breakthrough for the fully genetically-encoded strategy. In Engineering and Characterization of an Enhanced Fluorescent Protein Voltag Sensor (2007), The Knöpfel group tagged the recently discovered voltage sensitive phosphotase (Ci-VSP) with CFP and YFP FRET pairs in place of the phosphotase domain. This tagged protein expressed at the membrane much more efficiently than previous genetically encoded voltage sensors based on potassium channel subunits. By injecting physiological voltage changes and averaging 50-90 traces, they were able to pull out a few percent ratio change from a brief series of action potentials. Single spikes were resolvable. Although this sensor (VSFP2.1) was pretty slow (tau > 10ms), this new substrate looked promising for future sensor development.

They have since sped the response up. In Engineering of a Genetically Encodable Fluorescent Voltage Sensor Exploiting Fast Ci-VSP Voltage-Sensing Movements (2008 ), they determined that the gating motion of the voltage sensing component was very fast (~1ms), while the fluorescence change was slow (~100ms). So they did what any good FRET tinkerer would do, chop away at the linkers between FP components. The sensor response improved, and they noticed that there was a disconnect between the speed of the CFP and YFP responses. Not only was CFP decreasing from enhanced FRET, it was being directly quenched by interactions with the lipid membrane. Chopping off the YFP from the the construct then dramatically increased the speed of the CFP quench. This improved sensor, VSFP3.1 has an activation time constant of 1.3ms, though it’s response magnitude is still quite small (a few % dF/F).

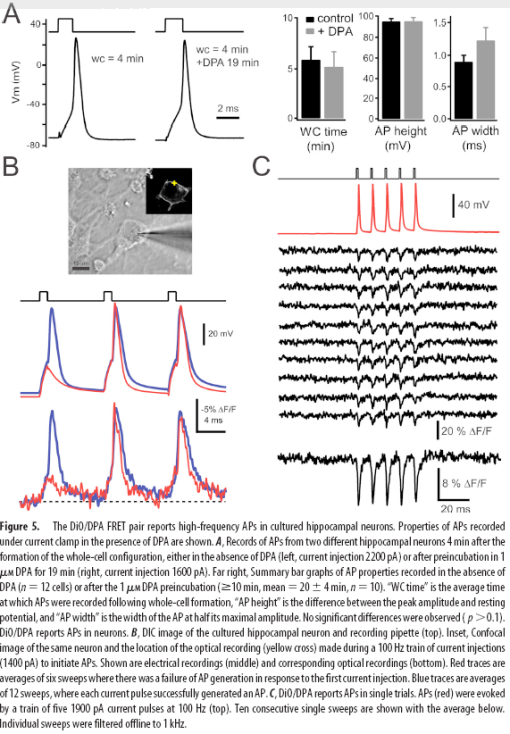
A hybrid approach to measuring electrical activity in genetically specified neurons (2005) has a much greater response magnitude. Pancho Bezanilla’s group exploited the rapid, voltage-dependent translocation of the small molecule quencher dipicrylamine (DPA) through the plasma membrane to change the fluorescence of membrane-teathered GFP in a voltage-dependent manner. Responses of the hybrid voltage sensor (hVOS) were relatively large (34% per 100mV) and fast (0.5ms). Single action potentials were detectable without averaging. However, since DPA is a charged molecule, it significantly increased the capacitance of the membrane. The levels of DPA required to see large responses inhibited action potentials and were intolerable to neurons.

Last month in Rational Optimization and Imaging In Vivo of a Genetically Encoded Optical Voltage Reporter (2008 ), Sjulson and Miesenböck reported optimized parameters for the hVOS approach. They built a quantitative model of the quenching effects of DPA on membrane-teathered GFP. The quenching is limited by the distance the DPA can approach the chromophore of GFP. Only the closest DPA molecule to the chromophore significantly contributes to a GFP’s quenching. After lots of pretty heat maps and graphs, the model tells them to chop off the tail of EGFP to bring the C-terminal tethering sequence closer to chromophore. I should note that an 11 amino acid C-terminal truncation of ECFP has improved the response of a tremendous number of FRET reporters and has been standard practice for the last 8 years. By shortening the linker they manage to triple the response size. I’d suggest, if they haven’t already, to lop off another six amino acids (end the EGFP with …LEFVTAA) and see if works. EGFP and ECFP usually tolerate it.

Using this optimized reporter, they are able to reduce DPA concentrations to levels that are usable in vivo, at least for a few minutes. They record fast optical responses to electrical activity in the Drosophila antennal lobe using 2uM DPA. But after a few minutes, the DPA loaded neurons become strongly inhibited.

The bottom line? Voltage-sensitive imaging has seen big progress in the last few years, but still has a long way to go to gently record single APs in a dish or in vivo. Or does it? I’m hearing whispers that a different group has developed a synthetic dye technique that is getting >10% dF/F to single APs with millisecond response times. Is it the real deal? Watch this space…






Recent Comments