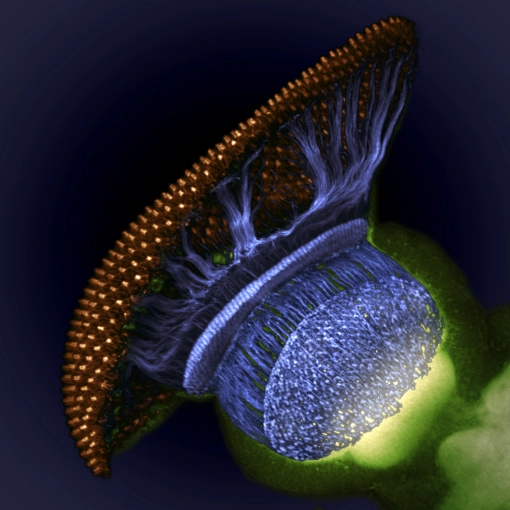
By twitter request, click-thru for the original resolution of a fellow Janelian’s Nikon Small World 4th place prize winner. Ryan Williamson composed the image.
Drosophila visual system
24 10 2012Comments : 1 Comment »
Tags: drosophila, Imaging, rentia
Categories : Imaging
Quick Picks : Brainbow flies
8 02 2011Nature methods published two papers which extend brainbow-like techniques of stochastic multicolored neuronal labeling into fruit flies. Nature’s summary explains the two methods.
The first technique, called dBrainbow, was developed by Julie Simpson, a neuroscientist at the Howard Hughes Medical Institute’s Janelia Farm Research Campus in Ashburn, Virginia, and her colleagues2. This method uses enzymes called recombinases to randomly delete some of the colour-producing genes from the string, leaving different genes next to the promoter regions in different cells. Individual cells are therefore uniquely coloured and so can be easily distinguished…
The second technique, called Flybow, was developed by Salecker and her colleagues3. They used an enzyme that ‘flips’ pairs of colour-producing genes on the string, leaving different genes next to the promoter region. The ‘flipping’ enzyme is also a recombinase, and so after being inverted, some of the colour-producing genes are randomly deleted. This ensures that all the different genes on the string can potentially end up next to the promoter, and be displayed by individual modified neurons.. Flybow uses a single string of four colours — red, green, blue and yellow.
These techniques will find use in building the structural and functional connectome of the fly.
Comments : 2 Comments »
Tags: brainbow, connectome, flybow, Imaging
Categories : genetics, Imaging, in vivo
UPDATE : DIADEM Final Results
15 09 2010The DIADEM automated neuronal reconstruction contest has finished. Accurate, fast, and high-resolution automated neuron reconstruction is of vital importance to cracking the mystery of how neural circuits perform. Even with perfect knowledge of the firing patterns of every cell in a circuit, our understanding of how these patterns are produced and how the information is processed would be quite limited. True understanding requires knowledge of the precise wiring diagram. This prize is a good first step towards bringing awareness of this tricky problem to the world’s best computer scientists.
$75,000 in prize money was to go to the group that was able to produce high-quality reconstructions of neuronal structures at least 20x faster than by-hand reconstructions. In the finals, the fastest speed achieved was 10X the by-hand method. Some groups were hindered by slight variances in the source data formatting, which normally isn’t a big deal unless you only have 20 minutes to produce as much reconstruction as possible…
Since no group was able to beat the hard floor, but substantial progress was made, the money was distributed amongst these finalists.
Badrinath Roysam Team, $25,000
“for the better overall generality of their program in producing robust reconstructions by integration of human and machines interactions.”
Armen Stepanyants Team, $25,000
“for the better overall biological results in the spirit of pure automation.”
Eugene Myers Team, $15,000
“for the excellent quality and strength of their algorithm.”
German Gonzalez Team, $10,000
“for their deeper potential, more original approach, and ultimate scalability of their proposed solution.”
Deniz Erdogmus Team
“for elevating themselves above the current state of automated reconstructions…with a deep understanding of the technical and scientific problems.”
Congrats to the placing teams.
Comments : 1 Comment »
Tags: circuit reconstruction, neuroanatomy, Neuroscience
Categories : brain imaging, Electron Microscopy, fluorescence, Imaging, in vitro, in vivo, neuroanatomy
Software Update : Ephus, ScanImage & Neuroptikon
20 08 2010Three excellent pieces of neuroscience software have been recently updated or freshly released. I have used two of them, Ephus and ScanImage, on a daily basis as primary data collection tools. The third, Neuroptikon, is quite useful for post-hoc illustration of neural circuits.

Ephus is a modular Matlab-based electrophysiology program that can control and record many channels of tools and data simultaneously. Under control of a sophisticated internal looper or external trigger, you can initiate an ephys recording, trigger camera frames, adjust galvo positions, open/close shutters, trigger optical stimulation, punishments, rewards, etc. It is a workhorse program for non-imaging related in vitro and in vivo electrophysiology experiments. Ephus is named for the fabled baseball pitch, and pronounced as “EFF-ess”. As with the pitch, it may trick you at first, but eventually you’re sure to hit a home run. Of course, the name also evokes electrophysiology, which is the fundamental orientation of the project, be it optical or electrical.
Ephus 2.1.0 is a major release, and the only official version at this time. The software is fully described in a publication in Frontiers in Neuroscience. New features include unlimited recording time, with disk streaming, for applications such as EEGs and long traces during in-vivo behavior. A number of additional scripts for in-the-loop control have been added. New configuration/start-up files have been created, with a template to help get up and running quickly. This release also includes a number of bug fixes.
ScanImage is another Matlab-related software program that is used for optical imaging and stimulation of neurons in vitro and in vivo. It finds much use a control platform for 2-photon imaging, glutamate uncaging and laser-scanning photostimulation. An early incarnation is described in this paper by Pologruto, et al. It provides a lot of power right out of the box (bidirectional scanning @ 0.5ms/line, etc) and is easily extensible via custom user function plugins.
Neuroptikon is a sophisticated network visualization tool. It can build Van Essen-like diagrams of any circuit you like, but it is so much more. The direction of communication is animated, and subsets of regions and connections can be brought into focus, which greatly eases the clarity of the network. The diagrams can be built in three-dimensions, to preserve relative topography, or functional grouping. There is simple GUI-based control, while more complex tasks can use a scripting interface. This is great software for anyone who needs to imagine information flow in a complex network.
All three tools are released for free use under the HHMI/Janelia Farm open source license.
Download Here :
Comments : Leave a Comment »
Tags: electrophysiology, Imaging, matlab, software, two-photon
Categories : circuit mapping, Glutamate, Imaging, in vitro, in vivo, optogenetics, photoactivation, software, Visualization
Cameleon-Nanos : High Affinity GECIs
9 08 2010Takeharu Nagai’s lab has published in Nature Methods, Spontaneous network activity visualized by ultrasensitive Ca2+ indicators, yellow Cameleon-Nano, demonstrating a new set of calcium indicators based on yellow cameleon. Back when he was still Take-san, Take’s ability to churn out and manually screen hundreds of cameleon variants was impressive and inspiring. With high-throughput GECI pipelines now ramping up at Janelia, the idea of laboriously screening 200 variations on a theme (be it cameleons or GluSnFRs), seems a bit archaic. However, this paper is a good example of the progress that can still be made by understanding the needed sensor parameters and fiddling with the primary amino acid structure in a relatively low-throughput way. Take-sensei’s results are another example of the pramatic rule in protein design, “when in doubt, tinker with the linker.”
The cameleon-nano family achieves greater apparent calcium affinity than YC2.60-4.60, reaching levels of up to 15nM. They did this by increasing the flexibility of the linker by extending the standard Tsien/Miyawaki/Baird Gly-Gly-Ser linker with additional glycines. In this case, the longer the linker between the CaM and M13 segments, the greater the apparent affinity. Interestingly, improvement by increasing linker flexibility is precisely the opposite the advice Atsushi and Take gave me for achieving high ratio changes with FRET reporters. Back at RIKEN in 2002, they suggested I use short, stiff linkers to restrict the rotational freedom of the fluorescent pairs. Then one could find orientations where relative rotation of dipole moment gave much greater FRET changes than would be expected from changes in FP distance alone. Take and Atsushi’s big YC2.60/3.60 paper strongly supported this idea! However, as our understanding of the ideal parameters of calcium sensor’s for in vivo imaging has grown, development directions have adjusted.
Cameleon-Nanos achieve higher signal/noise for sparse action potentials at the expense of linearity. Like Fluo-4, the signal saturates at relatively low AP frequencies. I think the absolute affinities measured for this family (15, 30, 50 and 140nM) should be considered very rough estimates. They extrapolated these values from stopped-flow binding experiments, because
Although we would like to measure the koff of YC2.60 and its high affinity variants such as YC-Nano15, we could not do it because it was very difficult to precisely control free Ca2+ concentration at around few tens of nM as far as we used EGTA (Kd for Ca2+ = 151 nM in 0.1 M ionic strength, pH 7.2 at 25 oC). For this purpose, much stronger Ca2+ chelator with a smaller Kd value was required. However there is no such Ca2+ chelator available now.
I’m not sure why they didn’t just use the higher affinity, Mg++ insensitive, chelator BAPTA to make the Kd measurements the right way, with a linear regression of log-log fluorescence/concentration values. Due to instrument dead time, and the high affinity, I didn’t like stopped-flow based Kd measurements in the early GCaMP papers, and I don’t like them now. Also, the apparent calcium Kd will be highly dependent on solution ionic strength and [Mg++] which is unreported. Despite these quibbles, which are important only inasmuch as they give insight into the mechanism of improvement and the direction of future development, the cameleon-nano family looks promising for mammalian brain imaging. I still wonder if, assuming the reported Kd values are relevant in vivo, YC2.60 would be the best of the bunch, since cortical neurons have a resting Kd of ~50nM, which implies that a single AP transient of say 200nM free [Ca++] increase would push the calcium levels right up into the sweet-spot of YC2.60’s sensitivity.
This is all the more interesting given the recent results in YC3.60 imaging from Maz Hasan’s group. Previously, he had shown that transgenic YC animals were pretty bad for imaging. However, AAV-mediated gene delivery of YC3.60 has significantly improved the responses of the YC family. I’m not sure if they are really up to GCaMP3 levels under identical in vivo conditions, but they might have better long-term protein stability (or that might depend on which viral serotype is used.) What about cameleon-nanos, what about YC2.60?
Comments : Leave a Comment »
Tags: cameleon, mutagenesis
Categories : Calcium, fluorescence, FRET, gfp, Imaging, in vitro, in vivo, Multiphoton, optogenetics
CNiFERS of Acetylcholine and Attention
10 03 2010“If you find yourself needing to reread this paragraph, perhaps it’s not that well written. Or it may be that you are low on acetylcholine.” Acetylcholine (ACh) is a major modulator of brain activity in vivo and its release strongly influences attention. If we could visualize when and where ACh is released, we could more fully understand the large trial to trial variance found in many in vivo recordings of spike activity, and perhaps correlate that to attentional and behavioral states mediated by ACh transmission.
Back in grad school, when I was desperately trying to figure out what biological question to answer with my GluSnFR glutamate sensor, I ended up in a meeting with Kleinfeld, his grad student Lee Schroder and Palmer Taylor. We plotted a strategy to make a FRET sensor for acetylcholine. Palmer had recently solved crystal structures of an acetylcholine binding protein bound to agonists and antagonists. Snails secrete this binding protein into their ACh synapses to modulate their potency. The structures showed a conformational change upon agonist binding. The hope was that by fusing CFP and YFP to the most translocated bits of the protein, they would be able to see an ACh dependent FRET change. I was skeptical that it would work, as the translocation was much less than with calmodulin-M13 or periplasmic binding proteins used in Cameleon and GluSnFR, but thought was at least worth a shot. FRET efficiency is highly dependent on dipole orientation, not just dipole distance, and you never know how a small conformational change might rearrange the FP dipoles…
Of course, the simple idea didn’t work. Instead of giving up on the first dozen attempts, they kept plugging away at alternative strategies for measuring ACh release, and eventually succeeded. In this Nature Neuroscience report, An in vivo biosensor for neurotransmitter release and in situ receptor activity, Nguyen et al demonstrate a mammalian cell based system for optically measuring ACh levels in an intact brain. They coexpressed M1 muscarinic receptors with the genetically-encoded calcium indicator TN-XXL in HEK293 cells. ACh binding to the M1 receptor induced IP3-mediated calcium influx. This calcium rise was then picked up by the TN-XXL and reported as a change in CFP/YFP fluorescence. The crazy part is that they took this cell culture assay and implanted the cells into the brains of living rats!
In culture, the response was highly sensitive and monotonic (for phasic response section, EC50 of 11 nM, a Hill coefficient of 1.9 and a maximum of ΔR/R = 1.1). In vivo, using two-photon imaging through a cortical window, they were able to see clear ACh responses in frontal cortex from electrical stimulation of the nucleus basalis magnocellularis, typically 200-μs current pulses of 200 μA @ 100Hz for 20-500ms.
This was essentially a in vivo proof of principal experiment, showing that one could image ACh release in spatially and temporally precise regions of the brain. However, the imaging was done under urethane anesthesia, which is a much different brain state than an awake, behaving animal. Are CNiFERs sensitive, powerful and stable enough to determine behavioral states via imaging in an awake animal? Would expressing GCaMP3 (an indicator with greater fluorescence dynamic range) improve the performance of the CNiFER system? We used a very similar assay with ACh applied to HEK cells during the initial screens for better GCaMPs. Or, is the performance more limited by the properties of the M1 receptor and the adapting nature of IP3-mediated calcium dynamics? CNiFERS provide an interesting platform for looking at ACh and potentially other G-protein mediated signaling, but it remains to be seen if labs that aren’t as technically proficient with two-photon rig will find it more useful than cyclic voltammetry for measuring acetylcholine levels.
Nature Neuroscience, 13 (1), 127-132 DOI: 10.1038/nn.2469![]()
Nguyen, Q., Schroeder, L., Mank, M., Muller, A., Taylor, P., Griesbeck, O., & Kleinfeld, D. (2009). An in vivo biosensor for neurotransmitter release and in situ receptor activity
Comments : 1 Comment »
Tags: acetylcholine, CNiFERs, FRET, Imaging, TN-XXL
Categories : brain imaging, Fluorescent Protein, FRET, Imaging, in vitro, in vivo
Compressed Sensing in Neuroscience
1 03 2010Wired has a nice lay-person write-up of the rapidly developing field of compressed sensing. This is a technique that allows accurate reconstructions of highly undersampled sparse datasets. This field really took off in 2004 when Emmanuel J. Candès discovered that a tomography phantom image could be reconstructed exactly even with data deemed insufficient by the Nyquist-Shannon criterion. It is probably the hottest topic in imaging theory today.

Modified Shepp-Logan phantom with enhanced contrast for visual perception.
According to this review, Compressed Sensing MRI, its successful application requires three conditions to be met :
- Transform Sparsity: The desired image must have a sparse representation in a known transform domain (i.e., it must be compressible by transform coding),
- Incoherence of Undersampling Artifacts: The aliasing artifacts in a linear reconstruction caused by k-space undersampling must be incoherent (noise-like) in the sparsifying transform domain.
- Nonlinear Reconstruction: The image must be reconstructed by a non-linear method which enforces both sparsity of the image representation and consistency of the reconstruction with the acquired samples.
These conditions are well met by MRI imaging. This decoding technique dramatically shortens the required sampling times in an MRI magnet, which reduces the impact of motion artifacts, the bane of high-resolution MRI.
Unfortunately, I don’t think it is very applicable to situations where signal/noise of the underlying source is poor, like counting action potentials in shot-noise limited in vivo calcium imaging. But it’s use is spreading into other related problems, such as mapping the functional connectivity of neural circuitry. Tao Hu and Mitya Chklovvskii apply the compressed sensing algorithms in Reconstruction of Sparse Circuits Using Multi-neuronal Excitation (RESCUME) from the latest Advances in Neural Information Processing Systems journal. They measure a post-synaptic neuron’s voltage while stimulating sequentially random subsets of multiple potentially pre-synaptic neurons. The sparseness of connectivity allows them to map connectivity much faster than by a sequential method.
UPDATE: If you want to get a better sense of the breadth and depth of the applications of compressed sensing, check out Igor Carron’s comprehensive site, Compressive Sensing : The Big Picture.
Comments : 2 Comments »
Tags: algorithms, compressed sensing, connectomics, mri
Categories : circuit mapping, data analysis, Electron Microscopy, fMRI, Imaging, in vivo
Journal Club – In Vivo Inhibition Dynamics
18 02 2010Inhibition has a powerful role shaping the network dynamics of the cortex, but most studies of inhibitory circuitry are done in brain slice or anesthetized animals. In Membrane potential dynamics of GABAergic neurons in barrel cortex of behaving mice, Gentet et al use two-photon imaging to guide dual, whole-cell patch clamp of inhibitory and excitatory neurons in the mouse barrel cortex. These mice are head fixed, but awake and naturally whisking. The authors can then see how the membrane dynamics of both subthreshold and suprathreshold voltages are correlated across pairs of cells. Differences between the correlations for excitatory and inhibitory neurons shed light on how cortical circuitry processes sensory information in natural brain states.
For Journal Club #5, Mac Hooks, a post-doc here at Janelia working with Gordon Shepard and Karel Svoboda, walks us through these results. Also, there is a video introduction of the work by the lab head of the paper, Carl Petersen, provided by Cell Press.
Gentet, L., Avermann, M., Matyas, F., Staiger, J., & Petersen, C. (2010). Membrane Potential Dynamics of GABAergic Neurons in the Barrel Cortex of Behaving Mice Neuron, 65 (3), 422-435 DOI: 10.1016/j.neuron.2010.01.006
Comments : Leave a Comment »
Tags: in vivo, inhibition, patch clamp, two-photon, Voltage, whole cell
Categories : Imaging, in vivo, Voltage
Monte Carlo Calcium Spike Detection
9 02 2010I somehow missed that Josh Vogelstein’s method on action potential detection was published last summer. In Spike Inference from Calcium Imaging Using Sequential Monte Carlo Methods, the authors use a Monte Carlo approach to determine spike times from calcium imaging with superior performance to other deconvolution methods. It does a great job on simulated and in vitro data, I’d love to see performance on real in vivo recordings. If you are serious about calcium imaging, you should definitely get in touch with Josh and see what magic he can do with all that math. You should also ask him about the benefits of linen pants vs. denim, he’s got strong opinions on that subject as well…
Comments : Leave a Comment »
Tags: Calcium, deconvolution, Imaging, monte carlo
Categories : Calcium, data analysis, Imaging
Adaptive Optics for In Vivo Microscopy
13 01 2010Imaging fluorescence in an intact, living brain is difficult due to absorption and scattering of excitation and emission light. Two photon microscopy uses excitation light in the narrow optical window (700-950nm) where water and hemoglobin do not significantly absorb, which allows structure determination and functional imaging down to depths of ~600nm from the surface of the brain. However, scattering of the excitation light still occurs at these wavelengths, which distorts the excitation volume and causes a rapidly increasing fluorescent background at greater depths.

The vasculature was labeled by injecting flourescein dextran into the circulatory stream. The light source was a regenerative amplifier. ‘‘0 mm’’ corresponds to the top of the brain. Left, XZ projection. Right, examples of XY projections. Note the increase in background fluo- rescence deeper than 600 mm in the brain due to out-of-focus 2PE. (Theer et al., 2003)
In order to reduce this background and sharpen the two-photon excitation volume, Ji et al in Adaptive optics via pupil segmentation for high-resolution imaging in biological tissues adapt tricks from astronomy (see also Rueckel et al 2006). Depending on where each ray of excitation light enters the brain, the angles of scattering are different. By sequentially illuminating with spatially restricted subsections of full illumination beam, they see how the brain warps the light from each part of the beam. Then they use a flexible mirror and to change the angle and phase of the excitation beam so that each part of the beam comes together in the same place and phase. Since two-photon fluorescence scales as the square of the intensity of the excitation light, this dramatically improves both the resolution and the signal to noise of the fluorescence image.

Pre-warping the excitation light to cancel the scattering improves fluorescence localization and intensity.
How well will this work in the brain of a living animal? It’s not clear. Large differences in scattering paths across a field of view may dramatically slow the determination of optimal excitation beam warping and make it difficult to scan across a field of view quickly. Motion of the brain may change the pattern of scattering faster than the system can adapt. Still, the use of adaptive optics is one of the few promising techniques for increasing penetration depth and signal in optical brain imaging (not counting sucking off the bits of the brain that are “in the way” of the excitation beam.) There are other optical tricks in astronomy that have not yet been mined for neuroscience applications. Hopefully these will one day allow the functional imaging of all layers of the mammalian cortex, not just layer 2/3.
Comments : 3 Comments »
Tags: adaptive optics, two-photon microscopy
Categories : brain imaging, Imaging, in vitro, in vivo













Recent Comments