I somehow missed that Josh Vogelstein’s method on action potential detection was published last summer. In Spike Inference from Calcium Imaging Using Sequential Monte Carlo Methods, the authors use a Monte Carlo approach to determine spike times from calcium imaging with superior performance to other deconvolution methods. It does a great job on simulated and in vitro data, I’d love to see performance on real in vivo recordings. If you are serious about calcium imaging, you should definitely get in touch with Josh and see what magic he can do with all that math. You should also ask him about the benefits of linen pants vs. denim, he’s got strong opinions on that subject as well…
Monte Carlo Calcium Spike Detection
9 02 2010Comments : Leave a Comment »
Tags: Calcium, deconvolution, Imaging, monte carlo
Categories : Calcium, data analysis, Imaging
Three Cheers for GCaMP : Optogenetic Brain Reading
9 11 2009Three papers are out online in Nature Methods that show big improvements in calcium imaging with genetically encoded sensors. They are are based on the fluorescence intensity indicator, GCaMP. GCaMP, first developed by Junichi Nakai, consists of a GFP that has been circularly permuted so that the N and C termini are fused and new termini are made in the middle of the protein. Fused to one terminus is calmodulin and the other is a peptide, M13, that calmodulin (CaM) binds to in the presence of calcium. The name is supposed to look like GFP with a CaM inserted into it, G-CaM-P. Normally the GFP is dim, as there is a hole from the outside of its barrel into the chromophore. Upon binding calcium, this hole is plugged and fluorescence increases.

The first paper, A genetically encoded reporter of synaptic activity in vivo, from Leon Lagnado’s group, targets GCaMP2 to the outer surface of synaptic vesicles. This localization allows the fluorescence signal to be confined to the presynaptic terminal, where calcium fluxes in response to action potentials are high. This targeting improves the response magnitude of GCaMP2 and permits the optical recording of synaptic inputs into whatever region of the brain one looks at. They demonstrate the technique in live zebrafish.
In the second paper, Optical interrogation of neural circuits in Caenorhabditis elegans, from Sharad Ramanathan’s group, GCaMP2 has been combined with Channelrhodopsin-2 to perform functional circuit mapping in the worm. Since the worm’s structural wiring diagram has been essentially solved, functional data could say much about how “thick” the wires between each cell are. Unfortunately, with GCaMP2, the responses are too slow and weak to distinguish direct from indirect connections.
Finally, we have published a paper, Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators, describing the improved GCaMP3. This indicator has between 2-10x better signal to noise than GCaMP2, D3cpv and TN-XXL, depending on the system you are using. It’s kinetics are faster and it is more photostable than FRET indicators, and the responses are huge. When expressed in motor cortex of the mouse, neuronal activity is easily seen directly in the raw data. Furthermore, the sensor can be expressed stably for months, making it a potential tool for observing how learning reshapes the patterns of activity in the cortex.

Imaging of mouse motor cortex (M1) expressing the genetically-encoded calcium indicator GCaMP3 through a cortical window. After 72 days of GCaMP3 expression, large fluorescence transients can be seen in many neurons that are highly correlated with mouse running.
GCaMP3 is not perfect. It cannot reliably detect single action potential in vivo in mammals, though I doubt that any existing GECI can. Work continues on future generations of GCaMP that may achieve 100% fidelity in optical reading of the bits in the brain. However, there is considerable evidence from a number of groups that have been beta-testing the sensor, including the Tank lab of “quake mouse” fame, that it is a significant leap forward and unlocks much of the fantastic and fantasized potential of genetically-encoded calcium indicators.

Comparison of fluorescence changes in response to trains of action potentials in acute cortical slices.
I will try to post a more complete writeup of GCaMP3 for Brain Windows soon, with an unbiased eye to its strengths and weaknesses. We worked very hard to carefully characterize this sensor’s effects on cellular and circuit properties. If you have any questions about GCaMP3, please post them to the comments.
For further info about strategies for GECI use and optimization, check out our previous paper, Reporting neural activity with genetically encoded calcium indicators in Brain Cell Biology.
The official press release from HHMI regarding GCaMP3 is available here.
Comments : 26 Comments »
Tags: Calcium, GCaMP, GCaMP3, Neuroscience
Categories : brain imaging, Calcium, circuit mapping, fluorescence, Fluorescent Protein, in vitro, in vivo, neuroanatomy, Neuronal Control
Symposium : A Revolution in Fluorescence Imaging
11 02 2009
This coming Tuesday and Wednesday (Feb 17th & 18th) at UCSD, there will be a symposium honoring Roger Tsien, featuring presentations from 32 former and current members of the Tsien Lab. The topics are quite diverse, concentrated in genetically-encoded indicators, but also featuring fluorescent cell penetrating peptides for cancer therapy, photophore ligases for imaging synaptic development, and even a radical new design for the internal combustion engine.
The quality of speakers and subjects looks to be outstanding. Here is a complete schedule. You may notice that at 11:15 AM on Tuesday in Price Center East Ballroom, I will be presenting recent progress we have made in the development of genetically-encoded calcium indicators and their application to in vivo imaging. Don’t miss that one! 🙂 Roger’s talk, which will assuredly be equal parts absorbing, humorous, and illuminating, is at 4pm Wednesday in the Price Center Theater.
If you live in Southern California and are interesting in imaging technology, there isn’t a better place to be than this symposium. If you can’t make it, Brain Windows will have a full write-up following the event.
Here is the un-official schedule.
Tuesday February 17th – Price Center East Ballroom
9:00 -9:05 Varda Levram -Ellisman Opening
9:05-9:15 Palmer Taylor
Designing the next generation of genetically encoded sensors
9:15-9:30 Roger Heim
FRET for compound screening at Aurora/Vertex
9:30-9:45 Amy Palmer
Designing and using genetically encoded sensors: Lessons I learned from Roger
9:45-10:00 Robert Campbell
Beyond brightness: colony screens for fluorescent protein photo stability and biosensor FRET changes
10:00-10:15 Colette Dooley
GFP sensors for reactive oxygen species: Tying up loose ends and looking forward.
10:15-10:30 Peter Wang
Fluorescent Proteins and FRET biosensors for visualizing cell motility and mechanotransduction
Fluorescent proteins in neuroscience
11:00-11:15 Brian Bacskai
Aberrant calcium homeostasis in the Alzheimer mouse brain
11:15-11:30 Andrew Hires
Watching a mouse think: Novel fluorescent genetically-encoded calcium indicators applied to in vivo brain imaging
11:30-11:45 Alice Ting
Imaging synapse development with engineered photophore ligases
11:45-12:00 Rex Kerr
3D calcium imaging in C. elegans
Clinical applications
12:00-12:15 Todd Aguilera
Activatable Cell Penetrating Peptides for use in clinical contrast agent and therapeutic development
12:15-12:30 Quyen Nguyen
Surgery with Molecular Fluorescence Imaging Guidance
Fluorescent probes (Chemistry)
1:30-1:45 Tito Gonzalez
Voltage-Sensitive FRET Probes & Applications
1:45-2:00 Paul Negulescu
From watching ions to moving them
2:00-2:15 Timothy Dore
Roger-Inspired Photochemistry: Releasing Biological Effectors with 2PE
2:00-2:15 Joe Kao
Electron Paramagnetic Resonance Imaging in Living Animals
2:15-2:30 Brent Martin
Chemical probes for studying protein acylation
2:30-2:45 Jianghong Rao
Non-GFP based probes for imaging of the hydrolytic enzyme activity
Cellular research with and without Fluorescent probes
3:15-3:30 Carsten Schultz
Cell membrane repair visualized by GFP fusion proteins
3:30-3:45 David Green
Transcriptomes and Systems Biology: application to early mammalian embryogenesis
3:45-4:00 Clotilde Randriamampita
Paradoxical aspects of T cell activation revealed with fluorescent proteins
4:15-4:30 Wen-Hong Li
Studying dynamic cell-cell communication in vivo by Trojan-LAMP
4:30-4:45 Martin Poenie
Aim and Shoot: Two roles for dynein in T cell effector function
4:45-5:00 Gregor Zlokarnik
From bla to blah, blah in 20 years
5:00-5:15 James Sharp
President, Zeiss MicroImaging Gmbh
February 18 2009 – Leichtag 107
Cellular research with and without fluorescent proteins
9:00-9:15 David Zacharias
Fluorescent Proteins, Palmitoylation and Cancer: two out of three ain’t bad
9:15-9:30 Jin Zhang
Visualization of Cell Signaling Dynamics: A Tale of MAPK
9:30-9:45 Paul Sammak
Nuclear organization and movement in pluripotent stem cells measured by Histone GFP H2B
Branching out
9:45-10:00 Yong Yao
NIH Toolbox Program
10:00-10:15 Oded Tour
The Tour Engine – A novel Internal Combustion Engine with the potential to boost efficiency and cut emissions
Into the future
10:45-11:00 Xiaokun Shu
Visibly and infrared fluorescent proteins: photophysics and engineering
11:00-11:15 Michael Lin
Engineering fluorescent proteins for visualizing newly synthesized proteins and improving FRET-based biosensors
11:15-11:30 Jeremy Babendure
Training our next generation of Fluorescent Protein Enthusiasts
Main Event – Price Center Theater
4:00-5:00 Roger Tsien
Chancellor invitational lecture 2008 Nobel Prize in Chemistry
Comments : 6 Comments »
Tags: brain imaging, Calcium, Fluorescent Protein, FRET, GECI, gfp, Glutamate, Imaging, in vitro, Neuroscience, San Diego, UCSD
Categories : brain imaging, Calcium, Fluorescent Protein, FRET, gfp, Imaging, in vitro, in vivo, Meeting
Update : Structure of G-CaMP2
12 01 2009Today, Brain Windows welcomes its first guest contributor. Dr. Jasper Akerboom is a post-doctoral associate in the lab of Loren Looger at Janelia Farm, and is the lead author on a recently published report on the structure of the genetically-encoded calcium sensor, G-CaMP2. We are very grateful for his contribution!
After the previous post describing G-CaMP2 crystallization two papers describing the crystallization and structure determination appeared online:
Crystal structures of the GCaMP calcium sensor reveal the mechanism of fluorescence signal change and aid rational design. Akerboom J, Vélez Rivera JD, Rodríguez Guilbe MM, Alfaro Malavé EC, Hernandez HH, Tian L, Hires SA, Marvin JS, Looger LL, Schreiter ER. J Biol Chem. 2008 Dec 18.
and
Structural Basis for Calcium Sensing by GCaMP2. Wang Q, Shui B, Kotlikoff MI, Sondermann H.Structure. 2008 Dec 12;16(12):1817-27.
Both papers are very similar, with minor differences in the approach of some of the problems, which will be described below.
In the paper of Wang et al., crystallization of GCaMP2 is achieved by removing the pRSET tag, important for in vivo GCaMP2 function (Nakai et al). Removal of disordered expression tags often is essential for protein crystallization, however, Akerboom et al crystallized GCaMP2 with this tag still present. Spectrophotometric properties of purified GCaMP2 protein with and without pRSET module are identical.
Both in the JBC paper as well as the Structure paper the authors describe the presence of dimeric calcium loaded GCaMP2, appearing as a minor fraction during gel filtration analysis.

Size-exclusion trace of calcium loaded (blue line) and calcium free (red line) G-CaMP2
Akerboom et al initially only crystallized the dimeric form of GCaMP2. Attempts to crystallize monomeric GCaMP2 failed. Selection and mutagenesis of amino acids partaking in the dimer interface in GCaMP2 resulted in the subsequent crystallization of monomeric GCaMP2. Wang and coworkers were able to crystallize both forms without mutagenesis, although their GCAMP2 molecule had its pRSET module removed, indicating a potential role for the pRSET peptide in dimerization.
Both monomeric and the dimeric crystal forms described in both papers are essentially the same.
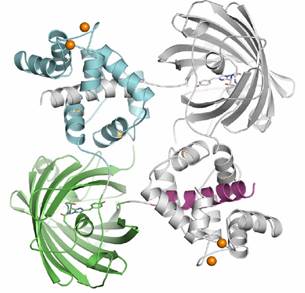
Dimeric G-CaMP2

Monomeric G-CaMP2
The dimeric form of G-CaMP2 is a domain swapped dimer with the M13 peptide (magenta) of each monomer bound by the calcium loaded CaM domain (cyan) of the other. The monomer is very different from the dimer, with the M13 peptide bound by the CaM domain of the own molecule. The interface between CaM and cpEGFP is considerably different between the two different oligomeric states of G-CaMP2.
Wang hypothesizes about a potential role of residue T116 (T203 in GFP numbering) playing in chromophore stabilization in calcium saturated G-CaMP2; this residue adopt a different rotamer in the dimeric structure, in a way that this threonine cannot partake in the hydrogen bond network, dimeric G-CaMP2 is less bright. In the paper by akerboom et al this residue adopts double conformations, so its not clear if this residue is actually the reason for this effect. In addition the mutation T203V results in increased fluorescence in G-CaMP2. Valine is hydrophobic and cannot participate in hydrogen bond formation at all.
Both groups performed mutational analysis of G-CaMP2. Both groups actually described a few identical positions (R81 and R377), and came roughly to the same conclusions, R81 and R377 play a role in the calcium loaded state of the protein. Wang et al performed the experiment using both mutations, and showed a profound decrease of fluorescence.
The group from Janelia Farm made some efforts to improve sensor functionality, and showed that replacing an aspartate close to the chromophore in the calcium saturated state with a tyrosine increases fluorescence by lowering the percentage of protonated chromophore.
Both Wang et al and Akerboom et al tried to study apo-G-CaMP2. Wang and co-workers used small-angle X-ray scattering (SAXS) of apo-G-CaMP2 and solved the structure of cpEGFP. The other group mutagenised all four EF-hands of CaM, removing the calcium binding capacity of G-CaMP2, and subsequently crystallizated the calcium binding deficient G-CaMP2. Both SAXS and crystallization indicated a more open structure of GCaMP2 compared to the calcium loaded state.


SAXS with fitted cpEGFP and 3CLN structures

apo G-CaMP2 structure
In the crystal structure, the M13 peptide and the C-terminal domain of CaM are disordered, indicating the large degree of freedom in apo G-CaMP2. Part of the linker between the M13 peptide and cpEGFP in the apo structure forms part of the beta barrel of cpEGFP.
Both papers will contribute to the understanding of the GECI G-CaMP2. Further directed mutagenesis studies on the basis of the results described in both manuscripts will hopefully result in a better sensor for in vivo imaging.
Comments : Leave a Comment »
Tags: Calcium, GECI, Imaging, structure
Categories : Calcium, Imaging, structure
BrainStorm 1 : The Calcium Memory Sensor
9 01 2009As mentioned in the previous post, this is the first installment of BrainStorm, a section of ideas I have under development, but don’t have the time to physically work on. This section will contain organically developed ideas, organized by project. Reader feedback is encouraged.
How can we identify the group of neurons that encode a particular thought?
I don’t want to simply see correlations of in activity of a few scattered neurons with a given thought, but identify the entire neuronal ensemble. Which neurons are active at a precise moment in a task? How are they wired together? Which are the drivers of activity?
Existing technology is inadequate to identify the entire neural ensemble that encodes a thought. Immediate early gene expression patterns have not been shown to be precisely correlated with brain activity, and have a temporal resolution on the order of minutes. Genetically encoded calcium sensors (GECIs) have the necessary temporal and spatial resolution, but their response is nearly as fleeting as a thought, making it impossible to simultaneously record from networks of thousands of possible participants with current microscopy techniques.
In BrainStorm 1, I will outline a technology, photoswitchable genetically-encoded calcium memory sensors, that can identify all the neurons in a large network that are active during user-specified, aribitrarly brief or long time periods. I will propose four potential strategies for construction of these sensors, and detail practical considerations for sensor design, screening and application.
Comments : Leave a Comment »
Tags: brain imaging, BrainStorm, Calcium, Fluorescent Protein
Categories : brain imaging, BrainStorm, Calcium, Fluorescent Protein, Imaging, in vivo, Multiphoton
Preview : Structure of G-CaMP2
10 12 2008A high-resolution crystal structure of the genetically-encoded calcium indicator G-CaMP2 would aid in rational design of improved calcium indicators. Crystallization of G-CaMP2 was first reported here :
Crystallization and preliminary X-ray characterization of the genetically encoded fluorescent calcium indicator protein GCaMP2
M. M. Rodríguez Guilbe, E. C. Alfaro Malavé, J. Akerboom, J. S. Marvin, L. L. Looger and E. R. Schreiter
Fluorescent proteins and their engineered variants have played an important role in the study of biology. The genetically encoded calcium-indicator protein GCaMP2 comprises a circularly permuted fluorescent protein coupled to the calcium-binding protein calmodulin and a calmodulin target peptide, M13, derived from the intracellular calmodulin target myosin light-chain kinase and has been used to image calcium transients in vivo. To aid rational efforts to engineer improved variants of GCaMP2, this protein was crystallized in the calcium-saturated form. X-ray diffraction data were collected to 2.0 Å resolution. The crystals belong to space group C2, with unit-cell parameters a = 126.1, b = 47.1, c = 68.8 Å, ![[beta]](https://i0.wp.com/scripts.iucr.org/logos/entities/beta_rmgif.gif) = 100.5° and one GCaMP2 molecule in the asymmetric unit. The structure was phased by molecular replacement and refinement is currently under way.
= 100.5° and one GCaMP2 molecule in the asymmetric unit. The structure was phased by molecular replacement and refinement is currently under way.
High-resolution atomic structures and mutational analysis were presented at SfN 2008 (see this previous post)
However, today a competing group has published an independent report on a similar set of G-CaMP2 structures in Cell Structure. More details to come…

Structural Basis for Calcium Sensing by GCaMP2
Qi Wang1, Bo Shui2,
Bo Shui2, Michael I. Kotlikoff2
Michael I. Kotlikoff2 and
and Holger Sondermann1,
Holger Sondermann1,
 ,
,


Genetically encoded Ca2+ indicators are important tools that enable the measurement of Ca2+ dynamics in a physiologically relevant context. GCaMP2, one of the most robust indicators, is a circularly permutated EGFP (cpEGFP)/M13/calmodulin (CaM) fusion protein that has been successfully used for studying Ca2+ fluxes in
the most robust indicators, is a circularly permutated EGFP (cpEGFP)/M13/calmodulin (CaM) fusion protein that has been successfully used for studying Ca2+ fluxes in vivo in the heart and vasculature of transgenic mice. Here we describe crystal structures of bright and dim states of GCaMP2 that reveal
vivo in the heart and vasculature of transgenic mice. Here we describe crystal structures of bright and dim states of GCaMP2 that reveal a sophisticated molecular mechanism for Ca2+ sensing. In the bright state, CaM stabilizes the fluorophore in an ionized state similar to that observed in EGFP. Mutational analysis confirmed critical interactions between the fluorophore and elements of the fused peptides. Solution scattering studies indicate that
a sophisticated molecular mechanism for Ca2+ sensing. In the bright state, CaM stabilizes the fluorophore in an ionized state similar to that observed in EGFP. Mutational analysis confirmed critical interactions between the fluorophore and elements of the fused peptides. Solution scattering studies indicate that the Ca2+-free form of GCaMP2 is a compact, predocked state, suggesting a molecular basis for the relatively rapid signaling kinetics reported for this indicator. These studies provide a structural basis for the rational design of improved Ca2+-sensitive probes.
the Ca2+-free form of GCaMP2 is a compact, predocked state, suggesting a molecular basis for the relatively rapid signaling kinetics reported for this indicator. These studies provide a structural basis for the rational design of improved Ca2+-sensitive probes.
Comments : 1 Comment »
Tags: Calcium, fluorescent, GECI, preview, structure
Categories : Calcium, Fluorescent Protein, gfp, Imaging, in vitro, Society for Neuroscience, structure
A brief history of calcium imaging
8 10 2008A few months ago I threw together a short presentation on the history of calcium imaging for a journal club here at Janelia. It is incomplete. It lacks notes. It is technical. It focuses much attention on early genetically-encoded indicators. However, calcium imaging is so intertwined with the work of Roger Tsien, my Ph.D. thesis advisor, and since he just won the Nobel Prize, I thought it might be of interest to some of the audience of Brain Windows. It does provide a little bit of background for some of the more recent developments chronicled on this site.
Enjoy.
Comments : 2 Comments »
Tags: Calcium, GECI, gfp, Imaging, Journal Club
Categories : Calcium, FRET, gfp, Imaging, Journal Club, Uncategorized
The great GECI shootout
21 07 2008Dierk Reiff’s lab has done another head-to-head in vivo showdown between various GECIs and a synthetic dye. Their paper, Fluorescence changes of genetic calcium indicators and OGB-1 correlated with neural activity and calcium in vivo and in vitro, is very interesting and deserves a full write-up. I will present a detailed analysis of the paper in a future update. For now, check the abstract.
Recent advance in the design of genetically encoded calcium indicators (GECIs) has further increased their potential fordirect measurements of activity in intact neural circuits. However, a quantitative analysis of their fluorescence changes (
F) in vivo and the relationship to the underlying neural activity and changes in intracellular calcium concentration (
[Ca2+]i) has not been given. We used two-photon microscopy, microinjection of synthetic Ca2+ dyes and in vivocalibration of Oregon-Green-BAPTA-1 (OGB-1) to estimate [Ca2+]i at rest and
[Ca2+]i at different action potential frequencies in presynaptic motoneuron boutons of transgenic Drosophila larvae. We calibrated
F of eight different GECIs in vivo to neural activity,
[Ca2+]i, and
F of purified GECI protein at similar
[Ca2+] in vitro. Yellow Cameleon 3.60 (YC3.60), YC2.60, D3cpv, and TN-XL exhibited twofold higher maximum
F compared with YC3.3 and TN-L15 in vivo. Maximum
F of GCaMP2 and GCaMP1.6 were almost identical. Small
[Ca2+]i were reported best by YC3.60, D3cpv, and YC2.60. The kinetics of
[Ca2+]i was massively distorted by all GECIs, with YC2.60 showing the slowest kinetics, whereas TN-XL exhibited the fastest decay. Single spikes were only reported by OGB-1; all GECIswere blind for
[Ca2+]i associated with single action potentials. YC3.60 and D3cpv tentatively reported spike doublets. In vivo, the KD(dissociation constant) of all GECIs was shifted toward lower values, the Hill coefficient was changed, and the maximum
F was reduced. The latter could be attributed to resting [Ca2+]i and the optical filters of the equipment. These results suggest increased sensitivity of new GECIs but still slow on rates for calcium binding.
Comments : Leave a Comment »
Tags: Calcium, Fluorescent Protein, FRET, Imaging, in vitro, in vivo
Categories : Calcium, Fluorescent Protein, FRET, Imaging, in vitro, in vivo
Pulse shaping for 2-photon signal enhancement
18 02 2008Gains in signal to noise ratios of organic dyes and genetically encoded indicators often come in modest steps following screening of large numbers of compounds or clones. Improvements are usually specific to individual chromophores, leading to the pigeonholing of development efforts on a small handful of indicators that have already undergone systemic optimization (i.e. cameleons, G-CaMP and troponin-based GECIs). Indicator photobleaching imposes strict limits on the amount of information which can be extracted by optical indicators. Improvement of specific indicators and their constituents is a worthy and necessary goal, but more generalizable improvements can be made by changing the nature of the illumination source. A series of papers from a variety of groups has shown that careful manipulation of the structure of pulse laser illumination can produce dramatic improvements in signal/noise and photobleaching during non-linear (two-photon) imaging. This is generalizable to numerous optical indicators. Reduction of photobleaching and photoinduced tissue damage will be essential for continuous optical monitoring of sparse neural activity.
My first encounter with these techniques came in 2003 during a lab presentation by Atsushi Miyawaki, who showed intriguing results with two-photon illumination of GFP. Kawano et al. shined ultra-short (28 femtosecond) pulses from a Ti-Sapphire laser on a plate of immobilized GFP. Due to the uncertainty principal, these ultra-short pulse durations cause a broad spread (~100nm) in the frequency of the laser pulse. They then actively modulated the phase of different frequency bands of the pulse. The interesting part is that they coupled this modulation to a feedback genetic algorithm that sought to increase the ratio of the GFP fluorescence to the intensity of the laser input. Over several hundred iterations of modulation, the system learned how to dramatically increase the output fluorescence over input power by tuning the phase of the frequency components. Using these optimally shaped pulses, they reduced the photobleaching rate by a factor of four! This was an impressive result, but it is unclear how useful this technique would be to live samples, in their heterogeneous aqueous environments. The tuning parameters might be less stable across a non-uniform sample.
The above paper raises intriguing questions on the nature of two-photon excited states and photobleaching. GFP and other fluorescent proteins have multiple bleaching modes, some permanent, some dark or UV-reversible. A more clear understanding of the photochemistry of bleaching could lead to improved illumination pulse design that could keep the chromophore away from these undesired states.
Early last year, Stenfan Hell’s group demonstrated a dramatic reduction in one and two-photon photobleaching by avoiding recurrent excitation of the GFP chromophore when it was in a dark absorbing state.Standard two-photon imaging procedure is to illuminate with a Ti-Sapphire laser pulsed at 80MHz, with a interpulse gap of 12.5ns. This gap is five times longer then the 2.4ns fluorescent lifetime of EGFP, giving the chromophore plenty of time to emit a photon and decay from the excited singlet S1 state. But is the singlet state the precursor to most photobleaching? Donnert et al. varied the pulse rate from 40 to 0.5MHz and discovered that photobleaching was dramatically reduced at the lower pulse rates, especially below 1MHz (1us interpulse interval). Under one photon illumination, total photons extracted from GFP before bleaching was increased 20-fold, while the rhodamine dye Atto532 increased by 8-fold. This suggests that the primary precursor to photobleaching is not the S1 state but is due to photon absorbtion during a dark triplet state T1, which has a relatively long lifetime of ~1us. Don’t illuminate during this state, and prevent most photobleaching! Under two photon illumination (800nm), even greater reductions in photobleaching of 25 and 20-fold respectively took place. This is particularly important because the much higher illumination power used in 2p excitation normally causes a dramatic, non-linear increase in the rate of photobleaching over 1p imaging in the region of focus.
What is special about the T1 triplet state that makes it more prone to causing photobleaching? Is it simply the longer lifetime gives a greater opportunity for an additional photon to hit, jumping to T2 and inducing a photochemical breakdown of the chromophore or the surrounding residues? Does T1 have a broader range of vibrational energies that can more easily engage the variety of photobleaching reactions than S1? How does the photobleaching rate of S2 compare to T2?
Despite the impressive reduction in photobleaching, and hence the greater S/N for a given bleach rate, Hell’s approach has a major drawback. Slowing the pulse train down ~100-fold also slows acquisition time down 100-fold. Therefore this technique is most useful for fixed specimens. Real-time, high resolution imaging of dynamic processes would be seriously degraded. There are work-arounds, such as wide-field pulsed illumination, rapid laser sweeping or multipoint parallel illumination, but these require additional technical development to make them feasible for the average, or even well-above average investigator.
Is there any related solution that can be easily applied to imaging live, dynamic cells? In this month’s Nature Methods, Na et al. from Eric Betzig’s group present an ‘exciting’ approach. They note that with conventional two photon illumination, most of the available laser power is wasted, intentionally blocked to reduce photodamage. As alluded to above, photodamage increases non-linearly with illumination intensity, making two-photon illumination methods particularly harmful. The authors demonstrate that this damage increases proportional to intensity to the ~2.4 power. To try to attenuate this effect, they used a series of mirrors to split the single ultra-short intense pulse (140 femtosecond) in half, and half again and again and again… They end up with 128 pulses of 1/128 the full intensity nearly evenly spaced every 37 picoseconds. This entire pulse group has a duration of 12ns, and with a 80MHz pulse cycle (12.5ns inter pulse interval) illuminates the chromophore with a steady stream of relatively dim pulsed light.
This split pulse illumination dramatically enhances acquisition speed and signal/noise. Images acquired at a rate of 0.4us/pixel with splitting look far more clear than those acquired at 25.6us/pixel without splitting. Photobleaching is reduced by a factor of four and acute photodamage is also reduced. Additional splitting may be possible and further improve the photobleaching attenuation. Importantly, they demonstrate this technique with GFP in fixed brain slices and in live worms and imagine dynamic responses with a calcium dye in living hippocampal slices. This technique appears to let you eat your cake and have it too. The implementation of the pulse splitting is modular, appears relatively simple to those with customizable two-photon instruments and works with existing Ti-Sapphire lasers. I anticipate rapid adoption by serious imaging labs.
Each of the above advances attacks the problem of indicator photobleaching by a different approach, and each focuses on a different aspect of the photochemistry. Theoretically, one could even combine all three for maximum photon collection efficiency before photobleaching, though this would also require combining the drawbacks of each. Photobleaching will continue to be a major concern in the imaging of dynamic processes, particularly when the signal is not synchronized with the onset of image acquisition. These techniques show substantial progress towards alleviating this concern, and I’m heartened to see a number of excellent labs are focusing so much energy on it.
A final question : How will each of these techniques affect acceptor photobleaching (I’m looking at you Citrine and Venus) in FRET imaging experiments? Do the same processes apply when the excitation is coming from a FRET donor?
Comments : 2 Comments »
Tags: Calcium, Fluorescent Protein, FRET, Multiphoton, Photobleaching
Categories : Calcium, Fluorescent Protein, FRET, Imaging, in vitro, in vivo, Multiphoton, Photobleaching
Three quick paper picks
18 10 2007Here are three papers that are worth reading over. No time for full reviews.
New Single-FP GECIs
The Russian fluorescent protein team has come out with some new single fluorescent protein G-CaMP/pericam-like sensors. They fiddled with the linker sites at the 145 and 148AA insertion points and found a great deal of fluorescence sensitivity to the amino acid composition at those sites. They note two new sensor constructs Case12 and Case16 that have 12-16.5x maximal changes in fluorescence upon calcium binding, a significant improvement over G-CaMP2. The tradeoff appears to be that they are dimmer. They show calcium responses in HeLa, PC-12 and cortical neuron cells, but no direct head-to-head with other sensors in cells.
Multipoint multiphoton microscopy
In this technical paper, an MIT group led by Peter So examines some issues surrounding multipoint excitation multiphoton microscopy. In theory, multipoint excitation will dramatically increase image acquisition rate through parallelization. However, this comes at the expense of large increases in background scattered light, which reduces optical resolution and penetration depth. They present an imaging system using multianode photomultiplier tubes that lets them acquire an 8×8 grid of multiphoton excitiation points. This technique, plus post-hoc deconvolution allows them to approach the resolution and depth of single point multiphoton systems, with a parallel array.
Effects of linker length and stiffness on FRET
This paper from Harold Erickson’s group carefully examines the role of linker length and stiffness in determining the amount of FRET transfer between CFP and YFP derivatives. They show that intrinsically unstructured domains of identical amino acid length produce significant differences in FRET between tethered FPs. They propose a formula for estimating the stiffness of linkers from the degree of FRET, which corroborates a 2006 study by Evers et al. They also demonstrate that the reported enhancement of FRET between the CyPet and YPet pair is due to an enhanced tendency for the proteins to dimerize. This reinforces my thinking that the most needed and generalizable improvement to two-FP FRET systems is an enhanced photostability of the FRET acceptor. Somebody please screen for a bleach-resistant YFP! Props to Michael Lin for pointing out the paper.
Comments : Leave a Comment »
Tags: Calcium, Fluoresecent Protein, FRET, Imaging, in vitro, Multiphoton
Categories : Calcium, Fluorescent Protein, FRET, Imaging, in vitro, Multiphoton





Recent Comments